Arioc Versions Save
Arioc: GPU-accelerated DNA short-read alignment
v1.51
1 year agoNotes for Arioc v1.51:
The current build is v1.51.3140 (17 May 2023).
This is a stable, performance-tested build.
Arioc v1.51 reference-genome lookup tables are incompatible with those generated by AriocE in release v1.43 and earlier. If you are upgrading to v1.51 from an earlier version of Arioc, please re-execute AriocE to create new lookup tables. Updated lookup tables for the human, mouse, and bread yeast reference genomes are available at ftp://ftp.ccb.jhu.edu/pub/data/Arioc, the FTP server for the Center for Computational Biology at Johns Hopkins University.
New features
- AriocU, AriocP: increased accuracy of MAPQ computation model
- AriocU, AriocP: configuration parameter
AtNis no longer supported - AriocU, AriocP: binary output formats specific for the Terabase Search Engine (
TSE,KMH) are no longer supported - AriocU, AriocP: in repetitive regions where the placement of a short variant (SNV, indel) is arbitrary (upper track in the following image), Arioc reports the "leftmost" location within the region (lower track in the following image); this behavior is consistent with BWA and Bowtie 2 and may facilitate variant identification in repetitive regions with low coverage:
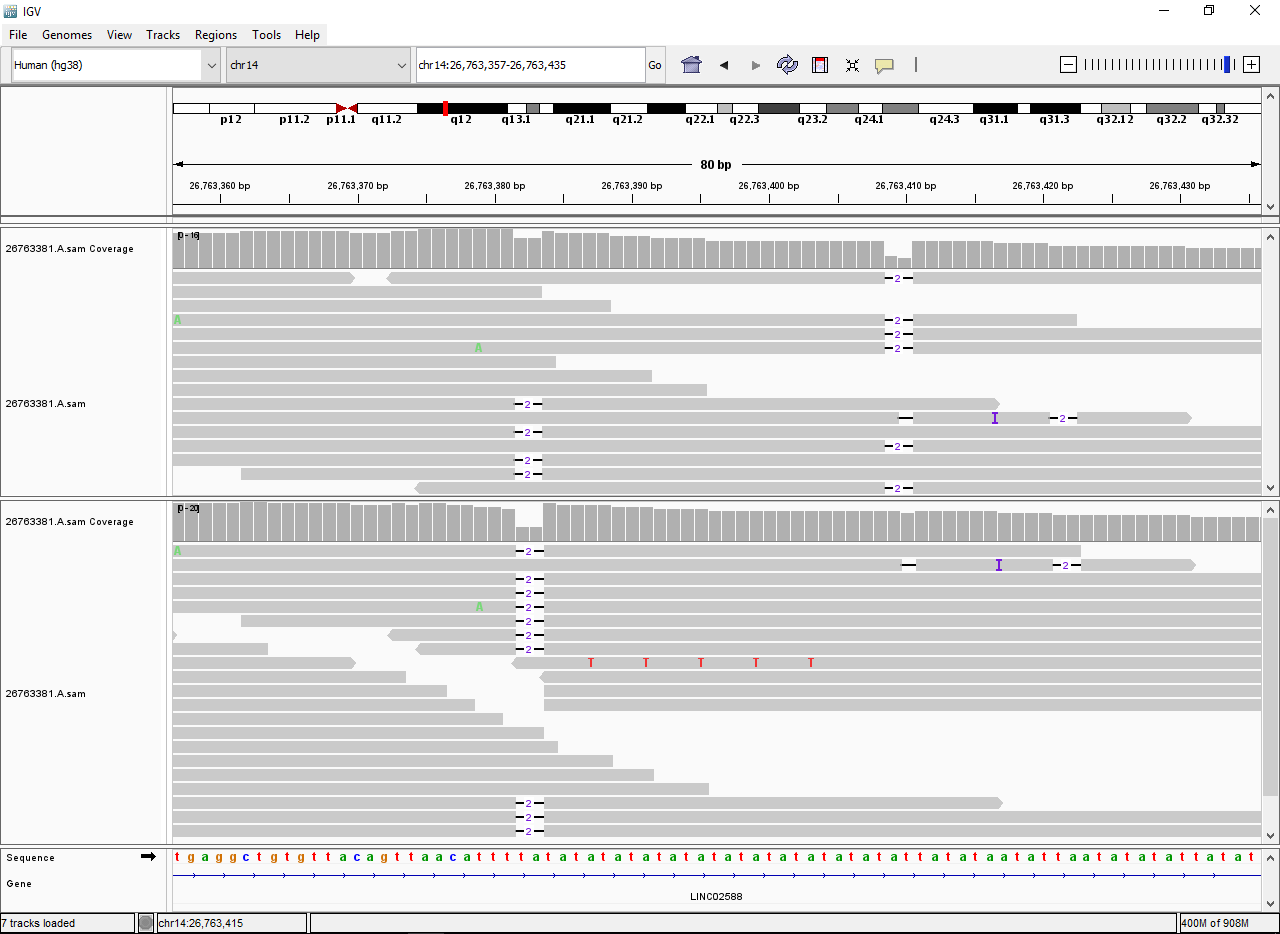
- AriocP: increased depth-of-search in gapped alignment of mates whose opposite mate has a reportable nongapped mapping
- AriocU, AriocP: in seed-and-extend gapped alignment, added a seed-prioritization heuristic that prioritizes higher-scoring mappings for the first seeds examined; this decreases (although it does not entirely eliminate) cases where a lower-scoring but reportable mapping precludes the identification of a higher-scoring mapping associated with the last seeds examined for a read sequence.
- AriocU, AriocP: added the
subRate("substitution rate") for tuning MAPQ performance; this replaces theerrorRateparameter implemented in previous Arioc versions. - AriocU, AriocP: added the
xmBQSparameter to specify a base quality score threshold for reporting methylation context for bisulfite-treated read sequences. - AriocU, AriocP:
maxJis now a required configuration parameter.
Bug fixes
- AriocE: corrupted defline in reference sequence longer than 21 million bases in FASTA file containing multiple reference sequences (see #31).
- AriocP: incorrect alignment score (AS) reported for occasional mates aligned by Smith Waterman alignment within regions "anchored" by their mapped opposite mates.
- AriocU, AriocP: data-dependent CUDA memory access exception when lookup tables are partitioned across four or more GPU devices (
useHJinGPmem="1"). - AriocE: possible error in multithreaded generation of genome configuration metadata.
Downloads
| file | content |
|---|---|
| Arioc.RQA.151.zip | Linux examples |
| Arioc.w.151.zip | Windows source code |
| Arioc.x.151.zip | Linux source code |
| AriocSetup.msi | Windows binaries and examples |
v1.50
2 years agoNotes for Arioc v1.50:
The current build is v1.50.3065 (10 February 2022).
This is a stable, performance-tested build.
Arioc v1.50 reference-genome lookup tables are incompatible with those generated by AriocE in release v1.43 and earlier. If you are upgrading to v1.50 from an earlier version of Arioc, please re-execute AriocE to create new lookup tables. Updated lookup tables for the human, mouse, and bread yeast reference genomes are available at ftp://ftp.ccb.jhu.edu/pub/data/Arioc, the FTP server for the Center for Computational Biology at Johns Hopkins University.
New features
- AriocE: support FASTA-formatted reference files that contain multiple individually-identified subunits (chromosomes, genomes, DNA regions)
- AriocE: support for SAM fields defined in FASTQ definition lines and subsequently appended to SAM alignment records
- XMC: added MAPQ, TLEN, and AS distributions
Bug fixes
- AriocU, AriocP: incorrect behavior with
suppressRaggSQ - AriocE: buffer overflow with very short reference genomes
- Linux and Windows examples: spurious
maxAparameter in AriocU and AriocP configuration files for GRCh38 - AriocU, AriocP: incorrect SEQ sporadically reported for reverse-complement reads whose length is an exact multiple of 21
Downloads
| file | content |
|---|---|
| Arioc.RQA.150.zip | Linux examples |
| Arioc.w.150.zip | Windows source code |
| Arioc.x.150.zip | Linux source code |
| AriocSetup.msi | Windows binaries and examples |
v1.43
3 years agoNotes for Arioc v1.43:
The current build is v1.43.2994 (19 April 2021).
New features
- AriocP: added
Vtwconfiguration parameter to relax minimum alignment score constraint on pairs where one mate is "hard to align" (see User Guide) - AriocU, AriocP: added
errorRateparameter for scaling MAPQ values (see User Guide) - AriocU, AriocP: refined MAPQ computation model to improve compatibility with GATK HaplotypeCaller
For comparison, here are some results from hap.py for whole genome sequencing of HG002 (PrecisionFDA Truth Challenge v2, "difficult-to-map regions"), GATK HaplotypeCaller v4.2.0, GIAB benchmark VCF and truthset confident regions, hard-filtered on QUAL, INFO.QD, and INFO.MQ using thresholds computed with RTG vcfeval:
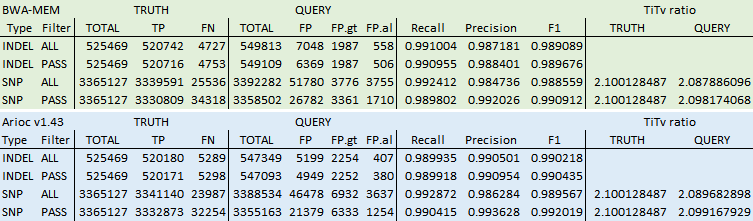
Bug fixes
- AriocP, AriocU: compile-time error with CUDA SDK v11.3.
Downloads
| file | content |
|---|---|
| Arioc.RQA.143.zip | Linux examples |
| Arioc.w.143.zip | Windows source code |
| Arioc.x.143.zip | Linux source code |
| AriocSetup.msi | Windows binaries and examples |
v1.42
3 years agoNotes for Arioc v1.42:
The current build is v1.42.2963 (30 November 2020).
New features
- AriocP: added TLEN distribution output as a space-separated list that can be copied and pasted for visualization or other analysis:
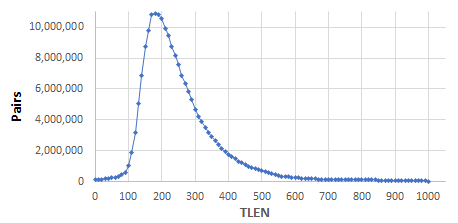
- AriocP: added performance metrics for TLEN skewness and fraction of discordant mappings due to out-of-range TLEN.
- AriocU, AriocP: added performance metrics for MAPQ distribution.
- AriocU, AriocP: added
xmClipparameter to control inclusion/exclusion of soft-clipped read sequence regions for BS-seq methylation context mapping (SAMXMfield). - AriocU, AriocP: implemented more stringent unduplication on reported "rejected" mappings.
Bug fixes
- AriocU, AriocP: incorrect POS reported for certain reads that straddle the origin of a reference sequence with circular topology
- AriocU, AriocP: incorrect XM reported for soft-clipped reads whose highest-scoring alignments map the reverse complement of the read sequence to the reverse complement of the reference ("non-directional" BS-seq alignments)
Downloads
| file | content |
|---|---|
| Arioc.RQA.142.zip | Linux examples |
| Arioc.w.142.zip | Windows source code |
| Arioc.x.142.zip | Linux source code |
| AriocSetup.msi | Windows binaries and examples |
v1.41
3 years agoNotes for Arioc v1.41:
The current build is v1.41.2923 (3 July 2020). This is a stable, performance-tested build.
New features
- MAPQ computed using exponentiated scores in a partition function
Bug fixes
(none)
Downloads
| file | content |
|---|---|
| Arioc.RQA.141.zip | Linux examples |
| Arioc.w.141.zip | Windows source code |
| Arioc.x.141.zip | Linux source code |
| AriocSetup.msi | Windows binaries and examples |
v1.40
4 years agoNotes for Arioc v1.40:
The current build is v1.40.2885 (1 May 2020).
This is a stable, performance-tested build.
Arioc v1.40 is not compatible with reference-genome lookup tables generated by release v1.30 and earlier. If you are upgrading to v1.40 from v1.30 or earlier, please re-execute AriocE to create new lookup tables as documented in the User Guide.
New features
- support for NVlink GPU memory interconnect
- more efficient concurrent use of available CPU threads
- added performance metrics (CUDA compute capability verification, GPU memory utilization, file output timing)
- support for circular reference-sequence topology
- added
AS(genome assembly identifier) andM5(MD5 checksum) tags to SAM@SQrecords - revised SAM
c3(triplet complexity score) tag to use DUST weighting and phred representation - reformatted tabular results to reconcile with output from samtools flagstat
- detailed discussion of performance tuning, including previously-undocumented extra (
X) parameters for fine-grained control
Bug fixes
- AriocP, AriocU: hang (infinite loop) with certain short QNAME fields
- AriocP: failure to recognize encoded query-sequence files (*a21.sbf) with certain multi-part filenames
- AriocE: segfault when encoding reads with certain multi-part filenames
- AriocU, AriocP: segfault when aligning samples that contain widely-varying read lengths
- AriocP: data-consistency validation exception with pairs where mate 1 has multiple nongapped mappings and mate 2 is unmapped
- AriocE: spurious error message at startup when no GPU is installed
- AriocU, AriocP: failure to emit CPU description in certain Power9 Linux configurations
- AriocU: duplicate mappings in SAM file with certain large-memory hardware configurations
- AriocP: inaccurate timing of GPU-to-CPU data-transfer performance metric for nongapped mappings
- AriocP, AriocU: updated SAM
NM(edit distance) computation to conform to current SAM optional fields specification
Downloads
| file | content |
|---|---|
| Arioc.RQA.140.zip | Linux examples |
| Arioc.w.140.zip | Windows source code |
| Arioc.x.140.zip | Linux source code |
| AriocSetup.msi | Windows binaries and examples |
v1.31
4 years agoNotes for Arioc v1.31:
The current build is v1.31.2566 (28 June 2019).
Arioc v1.31 is not compatible with reference-genome lookup tables generated by previous versions of the software. If you are upgrading to v1.31 from an earlier release, you must re-execute AriocE to create new lookup tables as documented in the User Guide.
New features
- user-specified reference sequence names (e.g., "chr1")
- new SAM tag c3 reports the number of distinct triplets in each read sequence
Bug fixes
(none)
Downloads
AriocSetup.msi: Windows binaries and examples Arioc.w.131.zip: Windows source code Arioc.x.131.zip: Linux source code Arioc.RQA.131.zip: Linux examples
v1.30
5 years agoNotes for Arioc v1.30:
The current build is v1.30.2559 (6 June 2019).
This is a stable, performance-tested build.
Arioc v1.30 is not compatible with reference-genome lookup tables generated by previous versions of the software. If you are upgrading to v1.30 from an earlier release, you must re-execute AriocE to create new lookup tables as documented in the User Guide.
New features
- maximum supported genome size is now 234 base pairs (17gbp)
- GPU-accelerated hashtable sorting in AriocE
- lookup table sizes are 5-10% smaller than in previous versions of Arioc
- optional MD string formatting: "standard" (strict conformance with SAM Format specification) or "compact" (saves space by omitting placeholder zeros between mismatched symbols, but incompatible with some software tools)
- improved parsing (and faster processing) of FASTQ defline and base quality score input
- minor changes to support Visual Studio 2019
Bug fixes
- AriocP: incorrect POS reported for a read whose mapping overlaps the start of the reference genome
- AriocE: unable to merge user-specified read group IDs
- AriocP, AriocU: fully-qualified executable file path not correctly displayed
- AriocP, AriocU: workaround for memory-allocation error handling with malloc() and realloc() in certain Linux implementations
- AriocE, AriocU, AriocP: insufficient buffer space for very long input filenames
- AriocP: divide-by-zero error in MAPQ computation for very short reads (~20 bases)
Downloads
AriocSetup.msi: Windows binaries and examples Arioc.w.130.zip: Windows source code Arioc.x.130.zip: Linux source code Arioc.RQA.130.zip: Linux examples
v1.25
5 years agoNotes for Arioc v1.25:
The current build is v1.25.2405.18255 (12 September 2018).
New features
- improved memory management of candidate reference-genome sequences in nongapped and gapped aligners
- progress messages (including estimated time remaining)
Bug fixes
- AriocU: data-dependent memory allocation error in gapped aligner
- AriocU, AriocP: intermittent thread-synchronization error (Linux version only)
- AriocU, AriocP: intermittent failure to correctly parse all attributes in the <A> element in configuration file
Downloads
AriocSetup.msi: Windows binaries and examples Arioc.w.125.zip: Windows source code Arioc.x.125.zip: Linux source code Arioc.RQA.125.zip: Linux examples
v1.24
6 years agoNotes for Arioc v1.24:
The current build is v1.24.2351.18110.
New features
- read-length distribution statistics in the AriocE (read encoding) component
- better memory management (larger batch sizes, more detailed error reporting) in the nongapped aligner component of AriocU and AriocP
Bug fixes
- AriocP: emit invalid FLAG bits for paired-end reads with one mate having exactly one mapping and the opposite mate having two or more mappings
- AriocU, AriocP: emit invalid XM (methylation context) map when a read's reverse complement maps to the reverse complement of the start of the reference sequence
- AriocP: emit incorrect MAPQ for paired-end mappings with dovetail orientation
Downloads
AriocSetup.msi: Windows binaries and examples Arioc.w.124.zip: Windows source code Arioc.x.124.zip: Linux source code Arioc.RQA.124.zip: Linux examples