Rumale Versions Save
Rumale is a machine learning library in Ruby
v0.22.5
3 years ago-
Added a new transformer that calculates the kernel matrix of given samples.
regressor = Rumale::Pipeline::Pipeline.new( steps: { ker: Rumale::Preprocessing::KernelCalculator.new(kernel: 'rbf', gamma: 0.5), krr: Rumale::KernelMachine::KernelRidge.new(reg_param: 1.0) } ) regressor.fit(x_train, y_train) res = regressor.predict(x_test) -
Added a new classifier based on kenelized ridge regression.
classifier = Rumale::KernelMachine::KernelRidgeClassifier.new(reg_param: 1.0) -
Nystroem now supports linear, polynomial, and sigmoid kernel functions.
nystroem = Rumale::KernelApproximation::Nystroem.new( kernel: 'poly', gamma: 1, degree: 3, coef: 8, n_components: 256 ) -
load_libsvm_file has a new parameter
n_featuresfor specifying the number of features.x, y = Rumale::Dataset.load_libsvm_file('mnist.t', n_features: 780) # p x.shape # => [10000, 780]
v0.22.4
3 years ago- Added classifier and regressor classes with voting-based ensemble method that combines estimators using majority voting.
require 'numo/openblas'
require 'parallel'
require 'rumale'
# ... Loading dataset
clf = Rumale::Ensemble::VotingClassifier.new(
estimators: {
log: Rumale::LinearModel::LogisticRegression.new(random_seed: 1),
rnd: Rumale::Ensemble::RandomForestClassifier.new(n_jobs: -1, random_seed: 1),
ext: Rumale::Ensemble::ExtraTreesClassifier.new(n_jobs: -1, random_seed: 1)
},
weights: {
log: 0.5,
rnd: 0.3,
ext: 0.2
},
voting: 'soft'
)
clf.fit(x, y)
v0.22.3
3 years ago- Added regressor class for non-negative least squares (NNLS) method. NNLS is a linear regression method that constrains non-negativeness to the coefficients.
require 'rumale'
rng = Random.new(1)
n_samples = 200
n_features = 100
# Prepare example data set.
x = Rumale::Utils.rand_normal([n_samples, n_features], rng)
coef = Rumale::Utils.rand_normal([n_features, 1], rng)
coef[coef.lt(0)] = 0.0 # Non-negative coefficients
noise = Rumale::Utils.rand_normal([n_samples, 1], rng)
y = x.dot(coef) + noise
# Split data set with holdout method.
x_train, x_test, y_train, y_test = Rumale::ModelSelection.train_test_split(x, y, test_size: 0.4, random_seed: 1)
# Fit non-negative least squares.
nnls = Rumale::LinearModel::NNLS.new(reg_param: 1e-4, random_seed: 1).fit(x_train, y_train)
puts(format("NNLS R2-Score: %.4f", nnls.score(x_test, y_test)))
# Fit ridge regression.
ridge = Rumale::LinearModel::Ridge.new(solver: 'lbfgs', reg_param: 1e-4, random_seed: 1).fit(x_train, y_train)
puts(format("Ridge R2-Score: %.4f", ridge.score(x_test, y_test)))
$ ruby nnls.rb
NNLS R2-Score: 0.9478
Ridge R2-Score: 0.8602
v0.22.2
3 years ago- Added classifier and regressor classes for stacked generalization that is a method for combining estimators to improve prediction accuracy:
require 'numo/openblas'
require 'parallel'
require 'rumale'
# ... Loading dataset
clf = Rumale::Ensemble::StackingClassifier.new(
estimators: {
rnd: Rumale::Ensemble::RandomForestClassifier.new(max_features: 4, n_jobs: -1, random_seed: 1),
ext: Rumale::Ensemble::ExtraTreesClassifier.new(max_features: 4, n_jobs: -1, random_seed: 1),
grd: Rumale::Ensemble::GradientBoostingClassifier.new(n_jobs: -1, random_seed: 1),
rdg: Rumale::LinearModel::LogisticRegression.new
},
meta_estimator: Rumale::LinearModel::LogisticRegression.new(reg_param: 1e2),
random_seed: 1
)
clf.fit(x, y)
v0.22.1
3 years ago- Add transfomer class for MLKR (Metric Learning for Kernel Regression) that performs transformation/projection along to the target variables. The following are examples of transforming toy example data using PCA and MLKR.
require 'rumale'
def make_regression(n_samples: 500, n_features: 10, n_informative: 4, n_targets: 1)
n_informative = [n_features, n_informative].min
rng = Random.new(42)
x = Rumale::Utils.rand_normal([n_samples, n_features], rng)
ground_truth = Numo::DFloat.zeros(n_features, n_targets)
ground_truth[0...n_informative, true] = 100 * Rumale::Utils.rand_uniform([n_informative, n_targets], rng)
y = x.dot(ground_truth)
y = y.flatten
rand_ids = Array(0...n_samples).shuffle(random: rng)
x = x[rand_ids, true].dup
y = y[rand_ids].dup
[x, y]
end
x, y = make_regression
pca = Rumale::Decomposition::PCA.new(n_components: 10)
z_pca = pca.fit_transform(x, y)
mlkr = Rumale::MetricLearning::MLKR.new(n_components: nil, init: 'pca')
z_mlkr = mlkr.fit_transform(x, y)
# After that, these results are visualized by multidimensional scaling.
PCA:
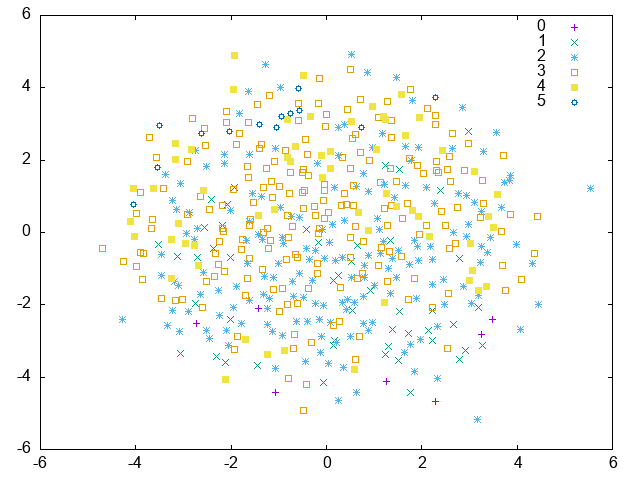
MLKR:
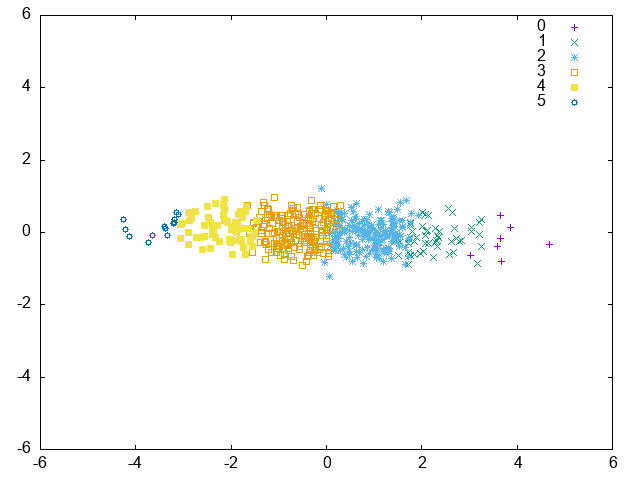
v0.22.0
3 years ago- Add lbfgsb.rb gem to runtime dependencies for optimization. This eliminates the need to require the mopti gem when using NeighbourhoodComponentAnalysis. Moreover, the lbfgs solver has been added to LogisticRegression, and the default solver changed from 'sgd' to 'lbfgs'. In many cases, the lbfgs solver is faster and more stable than the sgd solver.
v0.21.0
3 years ago- Change the default value of
max_iteronLinearModelestimators to 1000 from 200. The LinearModel estimators use stochastic gradient descent method for optimization. For convergence, it is better to set a large value for the number of iterations. Thus, Rumale has increased the default value of the max_iter.