Ludwig Versions Save
Low-code framework for building custom LLMs, neural networks, and other AI models
v0.7.1
1 year agoWhat's Changed
- Fixed confidence_penalty by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/3158
- Fixed set explanations by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/3163
- Bump to hummingbird 0.4.8 by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/3165
- Unpin pyarrow by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/3171
- Make Horovod an optional dependency when using Ray by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/3172
- Cherry-pick sample ratio changes by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/3176
- Fix TorchVision channel preprocessing (#3173) by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/3178
- Bump Ludwig to v0.7.1 (#3179) by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/3180
Full Changelog: https://github.com/ludwig-ai/ludwig/compare/v0.7...v0.7.1
v0.7
1 year agov0.7.beta
1 year agov0.6.4
1 year agoWhat's Changed
- Field fix: by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2714
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2719
- Bump Ludwig to 0.6.4 by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2720
Full Changelog: https://github.com/ludwig-ai/ludwig/compare/v0.6.3...v0.6.4
v0.6.3
1 year agoWhat's Changed
- Cherry-pick remote file syncing with hyperopt by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2644
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2646
- Cherry-pick bb8bef02c002eccbb6369292ac54490875bebbc4 by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2651
- Cherry-pick: Ensure no ghost ray instances are running in tests (#2607) by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2654
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2660
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2677
- Update version to v0.6.3 by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2682
Full Changelog: https://github.com/ludwig-ai/ludwig/compare/v0.6.2...v0.6.3
v0.6.2
1 year agoWhat's Changed
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2594
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2602
- 0.6.2: cherry-pick Explanation API by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2604
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2609
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2608
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2613
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2618
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2634
- Cherrypick: feat: adds max_batch_size to auto batch size functionality by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2632
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2636
- Update version to 0.6.2 by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2624
Full Changelog: https://github.com/ludwig-ai/ludwig/compare/v0.6.1...v0.6.2
v0.6.1
1 year agoWhat's Changed
- Cherry pick hyperopt plots by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2567
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2571
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2572
- fix: Limit frequency array to top_n_classes in F1 viz (#2565) by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2575
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2581
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2583
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2590
- Cherrypick: Comprehensive Configs by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2580
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2591
- Update version to 6.1 by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2582
- Readme fixes by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2592
Full Changelog: https://github.com/ludwig-ai/ludwig/compare/v0.6...v0.6.1
v0.6
1 year agoOverview
Ludwig 0.6 introduces several exciting features focused on modeling, deployment, and testing that make it more flexible, reliable, and easy to use in production.
- Gradient boosted models: Historically, Ludwig has been built around a single, flexible neural network architecture called ECD (for Encoder-Combiner-Decoder). With the release of 0.6 we are adding support for a different model architecture: gradient-boosted tree models (GBMs).
- Richer configuration schema and validation: We formalized the schema of Ludwig configurations and now validate it before initialization, which can help you avoid mistakes like typos and syntax errors.
- Probability calibration for binary and multi-class classification: With deep neural networks, the probabilities given by models often don't match the true likelihood of the data. Ludwig now supports temperature scaling calibration (On Calibration of Modern Neural Networks), which brings class probabilities closer to their true likelihoods in the validation set.
- Pipelined TorchScript: We improved the TorchScript model export functionality, making it easier than ever to train and deploy models for high performance inference.
- Model parameter update unit tests: The code to update parameters of deep neural networks can be too complex for developers to make sure the model parameters are updated. To address this difficulty and improve the robustness of our models, we implemented a reusable utility to ensure parameters are updated during one cycle of a forward-pass / backward-pass / optimizer step.
Additional improvements include a new global configuration section, time-based dataset splitting and more flexible hyperparameter optimization configurations. Read more about each specific feature below.
If you are learning about Ludwig for the first time, or if these new features are relevant and exciting to your research or application, we'd love to hear from you. Join our Ludwig Slack Community here.
Gradient Boosted Models (@jppgks)
Historically, Ludwig has been built around a single, flexible neural network architecture called ECD (for Encoder-Combiner-Decoder). With the release of 0.6 we are, adding support for a different model architecture: gradient-boosted tree models (GBM).
This is motivated by the fact that tree models still outperform neural networks on some tabular datasets, and the fact that tree models are generally less compute-intensive, making them a better choice for some applications. In Ludwig, users can now experiment with both neural and tree-based architectures within the same framework, taking advantage of all of the additional functionalities and conveniences that Ludwig offers like: preprocessing, hyperparameter optimization, integration with different backends (local, ray, horovod), and interoperability with different data sources (pandas, dask, modin).
How to use it
Install the tree extra package with pip install ludwig[tree]. After the installation, you can use the new gbm model type in the configuration. Ludwig will default to using the ECD architecture, which can be overridden as follows to use GBM:
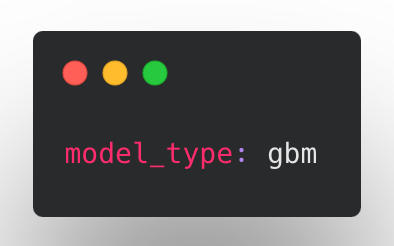
In some initial benchmarking we found that GBMs are particularly performant on smaller tabular datasets and can sometimes deal better with class imbalance compared to neural networks. Stay tuned for a more in-depth blogpost on the topic. Like the ECD neural networks, GBMs can be sensitive to hyperparameter values, and hyperparameter tuning is important to get a well-performing model.
Under the hood, Ludwig uses LightGBM for training gradient-boosted tree models, and the LightGBM trainer parameters can be configured in the trainer section of the configuration. For serving, the LightGBM model is converted to a PyTorch graph using Hummingbird for efficient evaluation and inference.
Limitations
Ludwig's initial support for GBM is limited to tabular data (binary, categorical and numeric features) with a single output feature target.
Calibrating probabilities for category and binary output features (@dantreiman)
Suppose your model outputs a class probability of 90%. Is there a 90% chance that the model prediction is correct? Do the probabilities given by your model match the true likelihood of the data? With deep neural networks, they often don't.
Drawing on the methods described in On Calibration of Modern Neural Networks (Chuan Guo, Geoff Pleiss, Yu Sun, Kilian Q. Weinberger), Ludwig now supports temperature scaling for binary and category output features. Temperature scaling brings a model's output probabilities closer to the true likelihood while preserving the same accuracy and top k predictions.
How to use Calibration
To enable calibration, add calibration: true to any binary or category output feature configuration:

With calibration enabled, Ludwig will find a scale factor (temperature) which will bring the class probabilities closer to their true likelihoods in the validation set. The calibration scale factor is determined in a short phase after training is complete. If no validation split is provided, the training set is used instead.
To visualize the effects of calibration in Ludwig, you can use Calibration Plots, which bin the data based on model probability and plot the model probability (X) versus observed (Y) for each bin (see code examples).
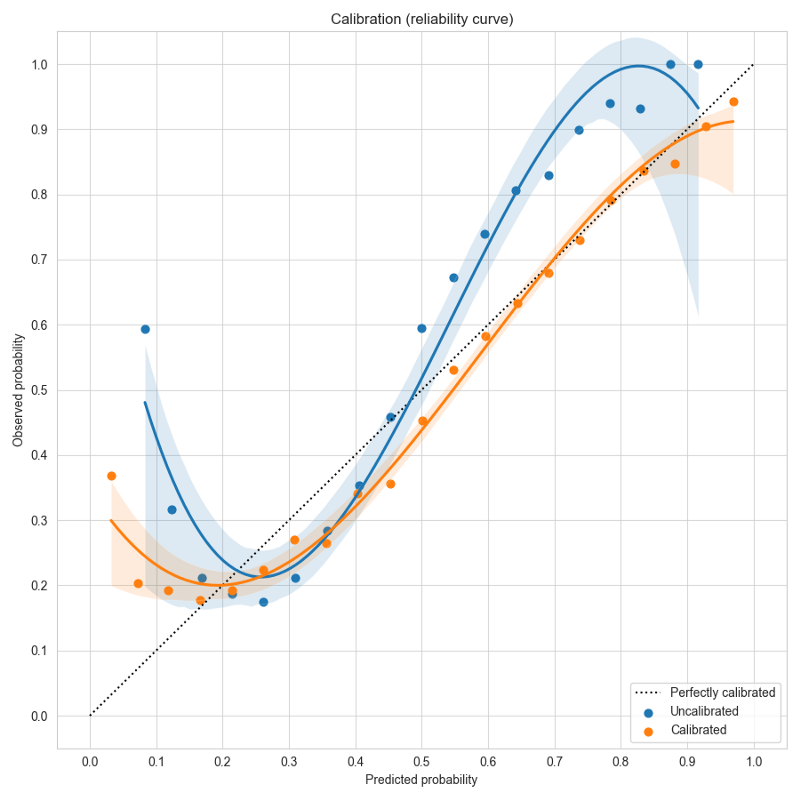
In a perfectly calibrated model, the observed probability equals the predicted probability, and all predictions will land on the dotted line y=x. In this example using the forest cover dataset, the uncalibrated model in blue gives over-confident predictions near the left and right edges close to probability values of 0 or 1. Temperature scaling learns a scale factor of 0.51 which improves the calibration curve in orange, moving it closer to y=x.
Limitations
Calibration is currently limited to models with binary and category output features.
Richer configuration schema and validation (@connor-mccorm @ksbrar @justinxzhao )
Ludwig configurations are flexible by design, as they internally map to Python function signatures. This allows configurations for expressive configurations with many parameters for the users to play with, but we have found that users would too easily have typos in their configs like incorrect value types or other syntactical inconsistencies that were not easy to catch.
We have now formalized the Ludwig config with a strongly typed schema, serving as a centralized source of truth for parameter documentation and config validation. Ludwig validation now explicitly restricts each parameter's values to valid ones, decreasing the chance of syntactical and logical errors and signaling immediately to the user where the issues lie, before processing data or starting training. Schemas also provide many future benefits including autocompletion.
Nested encoder and decoder parameters (@connor-mccorm )
We have also restructured the way that encoders and decoders are configured to now use a nested structure, consistent with other modules in Ludwig such as combiners and loss.
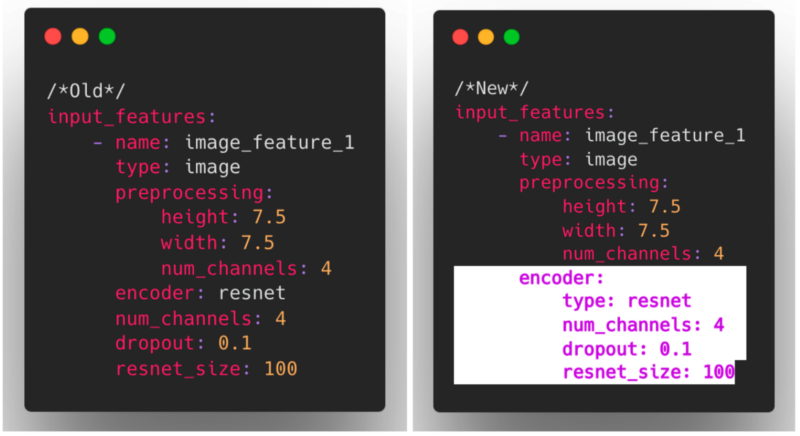
As these changes impact what constitutes a valid Ludwig config, we also introduced a mechanism for ensuring backward compatibility that invisibly and automatically upgrades older configs to the current config structure.
We hope with the new Ludwig schema and the improved encoder/decoder nesting structure, that you find using Ludwig to be a much more robust and user friendly experience!
New Defaults Ludwig Section (@arnavgarg1 )
In Ludwig 0.5, users could specify global preprocessing parameters on a per-feature-type basis through the preprocessing section in Ludwig configs. This is useful if users know they always want to apply certain transformations to their data for every feature of the same type. However, there was no equivalent mechanism for global encoder, decoder or loss related parameters.
For example, say we have a mammography dataset to predict breast cancer that contains many categorical features. In Ludwig 0.5, we might define our input features with encoder parameters in the following way:
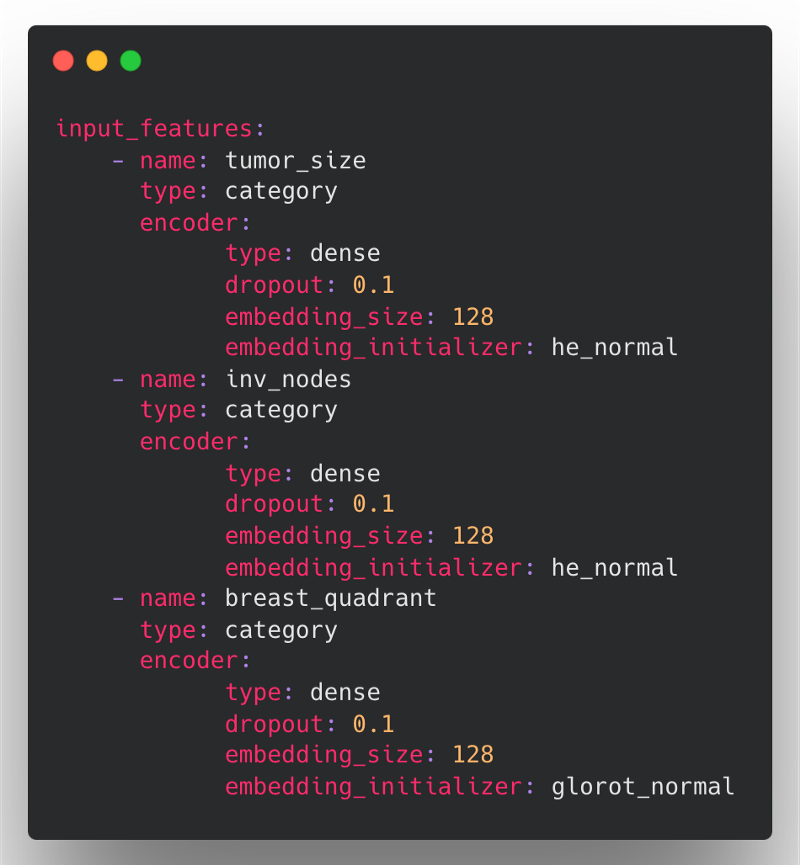
Here, the problem is that we have to redefine the same encoder parameters (type, dropout, and embedding_size) for each of the input features if we want to override the default value across all categorical features.
In Ludwig 0.6, we are introducing a new defaults section within the Ludwig config to define feature-type defaults for preprocessing, encoders, decoders, and loss. Default preprocessing and encoder configurations will be applied to all input_features of that feature type, while decoder and loss configurations will be applied to all output_features of that feature type.
Note that you can still specify feature specific parameters as usual, and these will override any default parameter values that come from the global defaults section.
The same mammography config above could be defined in the following, much more concise way in Ludwig 0.6:
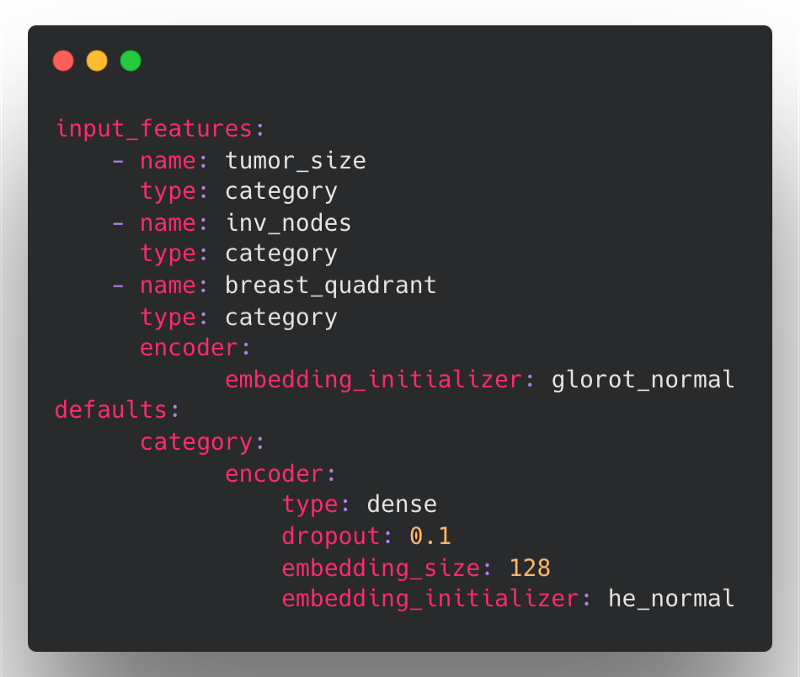
Here, the encoder defaults for type, dropout and embedding_size are applied to all three categorical features. The he_normal embedding initializer is only applied to tumor_size and inv_nodes since we didn't specify this parameter in their feature definitions, but breast_quadrant will use the glorot_normal initializer since it will override the value from the defaults section.
Additionally, in Ludwig 0.6, we have moved all global feature-type preprocessing within this new defaults section from the preprocessing section.
The defaults section enables the same fine-grained control with the benefit of making your config easier to define and read.
Global Defaults In Hyperopt (@arnavgarg1 )
The defaults section has also been added to hyperopt, so that users can define feature-type level parameters for individual trials. This makes the definition of the hyperopt search space more convenient, without the need to define individual parameters for each of the features in instances where the dataset has a large number of input or output features.
For example, if you want to hyperopt over different encoders for all text features for each of the trials, one can do so by defining a parameter this way:
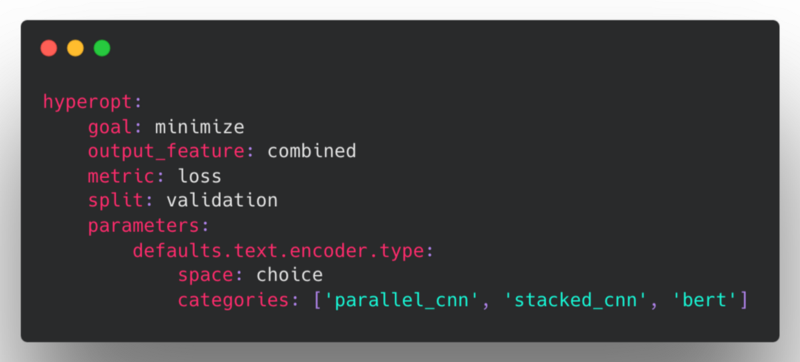
This will sample one of the three encoders for text features and apply it to all the text features for that particular trial.
Nested Configs In Hyperopt (@tgaddair )
We have extended the range of hyperopt parameters to support parameter choices that consist of partial or complete blocks of nested Ludwig config sections. This allows users to search over a set of Ludwig configs, as opposed to needing to specify config params individually and search over all combinations.
To provide a parameter that represents a full top-level Ludwig config, the . key name can be used.
For example, we can define a hyperopt search space where we sample partial Ludwig configs in the following way would create hyperopt samples that look like the following:
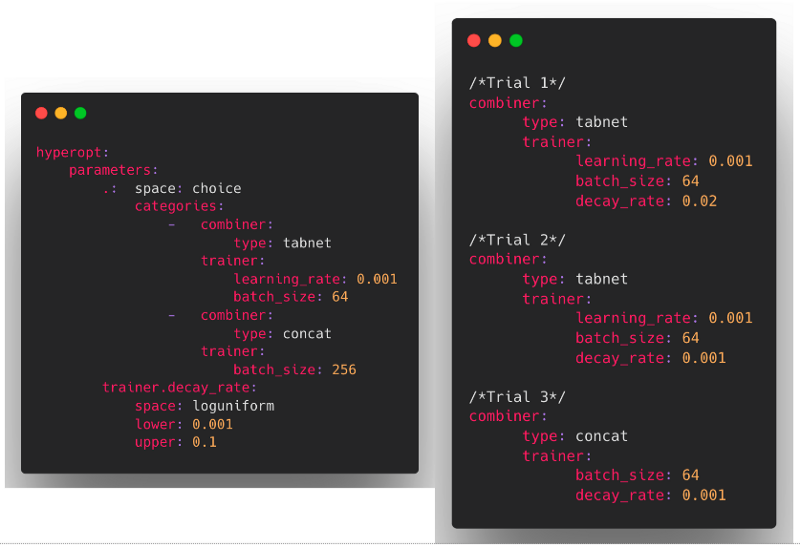
Pipelined TorchScript (@geoffreyangus @brightsparc )
In Ludwig v0.6, we improved the TorchScript model export functionality, making it easier than ever to train and deploy models for high performance inference.
At the core of our implementation is a pipeline-based approach to exporting models. After training a Ludwig model, users can run the export_torchscript command in the CLI, or call LudwigModel.save_torchscript. If model training was performed on a GPU device, doing so produces three new TorchScript artifacts:
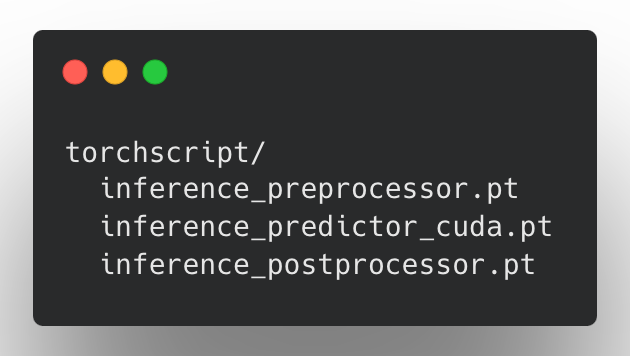
These artifacts represent a single LudwigModel as three modules, each separated by stage: preprocessing, prediction, and postprocessing. These artifacts can be pipelined together using the InferenceModule class method InferenceModule.from_directory, or with some tools such as NVIDIA Triton.
One of the most significant benefits is that TorchScripted models are backend and environment independent and different parts can run on different hardware to maximize throughput. They can be loaded up in either a C++ or Python backend, and in either, minimal dependencies are required to run model inference. Such characteristics ensure that the model itself is both highly portable and backward compatible.
Time-based Dataset Splitting (@tgaddair )
In Ludwig v0.6, we have added the ability to split based on a date column such that the data is ordered by date (ascending) and then split into train-validation-test along the time dimension. To make this possible, we have reworked the way splitting is handled in the Ludwig configuration to support a dedicated split section:
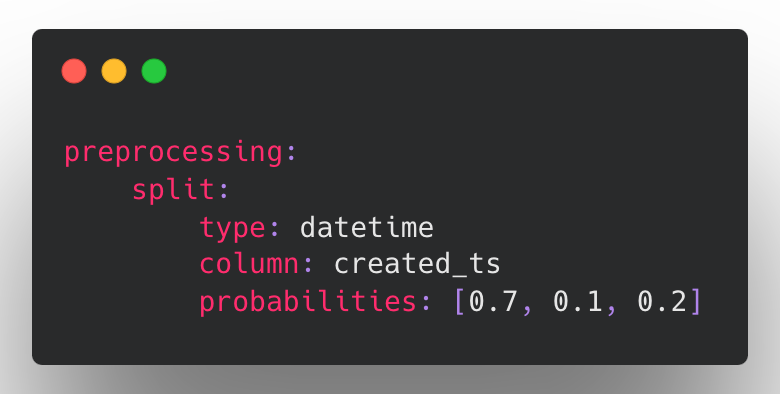
In this example, by setting probabilities: [0.7, 0.1, 0.2], the earliest 70% of the data will be used for training, the middle 10% used for validation, and the last 20% used for testing.
This feature is important to support backtesting strategies where the user needs to know if a model trained on historical data would have performed well on unseen future data. If we were to use a uniformly random split strategy in these cases, then the model performance may not reflect the model's ability to generalize well if the data distribution is subject to change over time. For example, imagine a model that is predicting housing prices. If we both train and test on data from around the same time, we may fool ourselves into believing our model has learned something fundamental about housing valuations when in reality it might just be basing its predictions on recent trends in the market (trends that will likely change once the model is put into production). Splitting the training from the test data along the time dimension is one way to avoid this false sense of confidence, by showing how well the model should do on unseen data from the future.
Prior to Ludwig v0.6, the preprocessing configuration supported splitting based on a split column, split probabilities (train-val-test), or stratified splitting based on a category, all of which were flattened into the top-level of the preprocessing section:
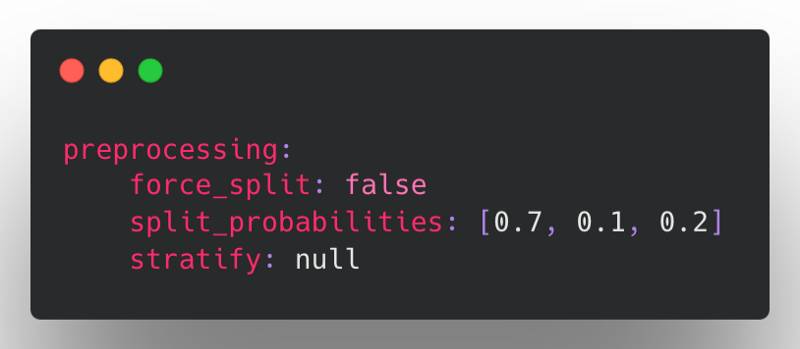
This approach was limiting in that every new split type required reconciling all of the above params and determining how they should interact with the new type. To resolve this complexity, all of the existing split types have been similarly reworked to follow the new structure supported for datetime splitting.
Examples
Splitting by row at random (default):
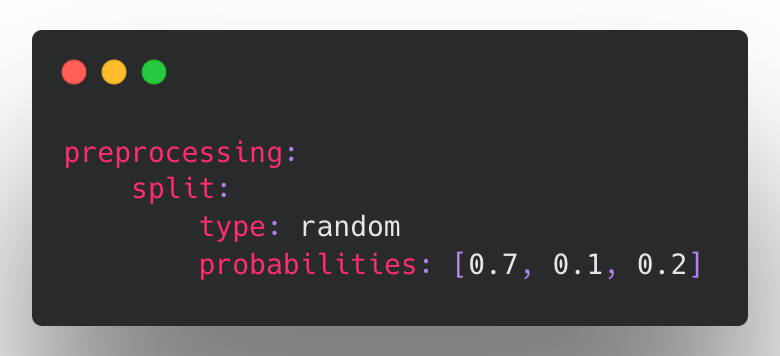
Splitting based on a fixed column.

Stratified splits using a chosen stratification category column.
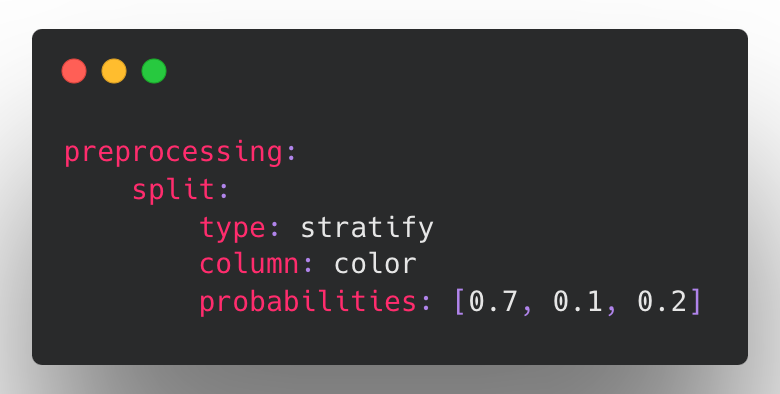
Be on the lookout as we continue to add additional split strategies in the future to support advanced usage such as bucketed backtesting. If you are interested in these kinds of scenarios, please reach out!
Parameter Update Unit Tests (@jimthompson5802 )
A significant step was taken in this release to improve the code quality of Ludwig components, e.g., encoders, combiners, and decoders. Deep neural networks have many layers composed of a large number of parameters that must be updated to converge to a solution. Depending on the particular algorithm, the code for updating parameters during training can be quite complex. As a result, it is near impossible for a developer to reason through an analysis that confirms model parameters are updated.
To address this difficulty, we implemented a reusable utility to perform a quick sanity check to ensure parameters, such as tensor weights and biases, are updated during one cycle of a forward-pass / backward-pass / optimizer step. This work was inspired by these earlier blog postings: How to unit test machine learning code and Testing Your PyTorch Models with Torcheck.
This utility was added to unit tests for existing Ludwig components. With this addition, unit tests for Ludwig now ensure the following:
- No run-time exceptions are raised
- Generated output are the correct data type and shape
- (New capability) Model parameters are updated as expected
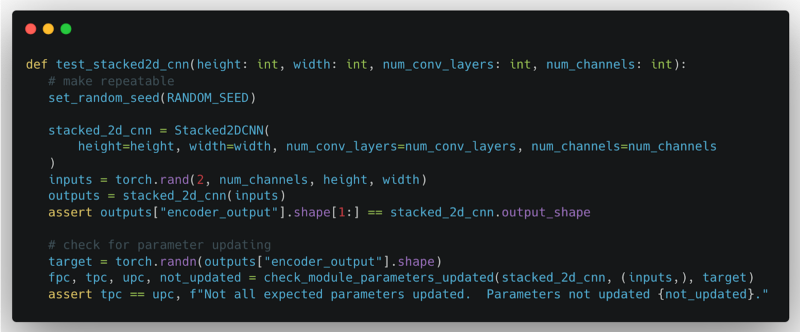
The above is an example of a unit test. First, it sets the random number seed to ensure repeatability. Next, the test instantiates the Ludwig component and processes synthetic data to ensure the component does not raise an error and that the output has the expected shape. Finally, the unit test checks if the parameters are updated under the different combinations of configuration settings.
In addition to the new parameter update check utility, Ludwig's Developer Guide contains instructions for using the utility. This allows an advanced user or a contributor, who is developing custom encoders, combiners, or decoders, to ensure the quality of their custom component.
Stay in the loop
Ludwig thriving open source community gathers on Slack, join it to get involved!
If you are interested in adopting Ludwig in the enterprise, check out Predibase, the declarative ML platform that connects with your data, manages the training, iteration, and deployment of your models, and makes them available for querying, reducing time to value of machine learning projects.
Full Changelog
- Fix ray nightly import by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2196
- Restructured split config and added datetime splitting by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2132
- enh: Implements
InferenceModuleas a pipelined module with separate preprocessor, predictor, and postprocessor modules by @brightsparc in https://github.com/ludwig-ai/ludwig/pull/2105 - Explicitly pass data credentials when reading binary files from a RayBackend by @jeffreyftang in https://github.com/ludwig-ai/ludwig/pull/2198
- MlflowCallback: do not end run on_trainer_train_teardown by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2201
- Fail hyperopt with full import error when Ray not installed by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2203
- Make convert_predictions() backend-aware by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2200
- feat: MVP for explanations using Integrated Gradients from captum by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2205
- [Torchscript] Adds GPU-enabled input types for Vector and Timeseries by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2197
- feat: Added model type GBM (LightGBM tree learner), as an alternative to ECD by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2027
- [Torchscript] Parallelized Text/Sequence Preprocessing by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2206
- feat: Adding feature type shared parameter capability for hyperopt by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2133
- Bump up version to 0.6.dev. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2209
- Define
FloatOrAutoandIntegerOrAutoschema fields, and use them. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2219 - Define a dataclass for parameter metadata. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2218
- Add explicit handling for zero-length image byte buffers to avoid cryptic errors by @jeffreyftang in https://github.com/ludwig-ai/ludwig/pull/2210
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2231
- Create dataset util to form repeatable train/vali/test split by @amholler in https://github.com/ludwig-ai/ludwig/pull/2159
- Bug fix: Use safe rename which works across filesystems when writing checkpoints by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2225
- Add parameter metadata to the trainer schema. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2224
- Add an explicit call to merge_wtih_defaults() when loading a config from a model directory. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2226
- Fixes flaky test test_datetime_split[dask] by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2232
- Fixes prediction saving for models with Set output by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2211
- Make ExpectedImpact JSON serializable by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2233
- standardised quotation marks, added missing word by @Marvjowa in https://github.com/ludwig-ai/ludwig/pull/2236
- Add boolean postprocessing to dataset type inference for automl by @magdyksaleh in https://github.com/ludwig-ai/ludwig/pull/2193
- Update get_repeatable_train_val_test_split to handle non-stratified split w/ no existing split by @amholler in https://github.com/ludwig-ai/ludwig/pull/2237
- Update R2 score to handle single sample computation by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2235
- Input/Output Feature Schema Refactor by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2147
- Fix nan in entmax loss and flaky sparsemax/entmax loss tests by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2238
- Fix preprocessing dataset split API backwards compatibility upgrade bug. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2239
- Removing duplicates in constants from recent PRs by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2240
- Add attention scores of the vit encoder as an additional return value by @Dennis-Rall in https://github.com/ludwig-ai/ludwig/pull/2192
- Unnest Audio Feature Preprocessing Config by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2242
- Fixed handling of invalud number values to treat as missing values by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2247
- Support saving numpy predictions to remote FS by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2245
- Use global constant for description.json by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2246
- Removed import warnings when LightGBM and Ray not requested by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2249
- Adds ability to read images from numpy files and numpy arrays by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2212
- Hyperopt steps per epoch not being computed correctly by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2175
- Fixed splitting when providing pre-split inputs by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2248
- Added Backwards Compatibility for Audio Feature Preprocessing by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2254
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2256
- Fix: Don't skip saving the model if the save path already exists. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2264
- Load best weights outside of finally block, since load may throw an exception by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2268
- Reduce number of distributed tests. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2270
- [WIP] Adds
inference_utils.pyby @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2213 - Run github checks for pushes and merges to *-stable. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2266
- Add ludwig logo and version to CLI help text. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2258
- Add hyperopt_statistics.json constant by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2276
- fix: Make
BaseTrainerConfigan abstract class by @ksbrar in https://github.com/ludwig-ai/ludwig/pull/2273 - [Torchscript] Adds
--deviceargument toexport_torchscriptCLI command by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2275 - Use pytest tmpdir fixture wherever temporary directories are used in tests. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2274
- adding configs used in benchmarking by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2263
- Fixes #2279 by @noahlh in https://github.com/ludwig-ai/ludwig/pull/2284
- adding hardware usage and software packages tracker by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2195
- benchmarking utils by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2260
- dataclasses for summarizing benchmarking results by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2261
- Benchmarking core by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2262
- Fixed default eval_batch_size when setting batch_size=auto by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2286
- Remove obsolete postprocess_inference_graph function. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2267
- [Torchscript] Adds BERT tokenizer + partial HF tokenizer support by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2272
- Support passing ground_truth as df for visualizations by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2281
- catching urllib3 exception by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2294
- Run pytest workflow on release branches. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2291
- Save checkpoint if train_steps is smaller than batcher's steps_per_epoch by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2298
- Fix typo in amazon review datasets: s/review_tile/review_title by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2300
- Refactor non-distributed automl utils into a separate directory. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2296
- Don't skip normalization in TabNet during inference on a single row. by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2299
- Fix error in postproc_predictions calculation in model.evaluate() by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2304
- Test for parameter updates in Ludwig components by @jimthompson5802 in https://github.com/ludwig-ai/ludwig/pull/2194
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2311
- Use warnings to suppress repeated logs for failed image reads by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2312
- Use ray dataset and drop type casting in binary_feature prediction post processing for speedup by @magdyksaleh in https://github.com/ludwig-ai/ludwig/pull/2293
- Add size_bytes to DatasetInfo and DataSource by @jeffreyftang in https://github.com/ludwig-ai/ludwig/pull/2306
- Fixes TensorDtype TypeError in Ray nightly by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2320
- Add configuration section for global feature parameters by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2208
- Ensures unit tests are deleting artifacts during teardown by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2310
- Fixes unit test that had empty Dask partitions after splitting by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2313
- Serve json numpy encoding by @jeffkinnison in https://github.com/ludwig-ai/ludwig/pull/2316
- fix: Mlflow config being injected in hyperopt config by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2321
- Update tests that use preprocessing to match new defaults config structure by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2323
- Bump test timeout to 60 minutes by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2325
- Set a default value for size_bytes in DatasetInfo by @jeffreyftang in https://github.com/ludwig-ai/ludwig/pull/2331
- Pin nightly versions to fix CI by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2327
- Log number of failed image reads by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2317
- Add test with encoder dependencies for global defaults by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2342
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2334
- Add wine quality notebook to demonstrate using config defaults by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2333
- fix: GBM tests failing after new release from upstream dependency by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2347
- fix: restore overwrite of eval_batch_size on GBM schema by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2345
- Removes empty partitions after dropping rows and splitting datasets by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2328
- fix: Properly serialize
ParameterMetadatato JSON by @ksbrar in https://github.com/ludwig-ai/ludwig/pull/2348 - Test for parameter updates in Ludwig Components - Part 2 by @jimthompson5802 in https://github.com/ludwig-ai/ludwig/pull/2252
- refactor: Replace bespoke marshmallow fields that accept multiple types with a new 'combinatorial'
OneOfFieldthat accepts other fields as arguments. by @ksbrar in https://github.com/ludwig-ai/ludwig/pull/2285 - Use Ray Datasets to read binary files in parallel by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2241
- typos: Update README.md by @andife in https://github.com/ludwig-ai/ludwig/pull/2358
- Respect the resource requests in RayPredictor by @magdyksaleh in https://github.com/ludwig-ai/ludwig/pull/2359
- Resource tracker threading by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2352
- Allow writing init_config results to remote filesystems by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2364
- Fixed export_mlflow command to not assume an existing registered_model_name by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2369
- fix: Fixes to serialization, and update to allow set repo location. by @brightsparc in https://github.com/ludwig-ai/ludwig/pull/2367
- Add amazon employee access challenge kaggle dataset by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2349
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2362
- Wrap read of cached training set metadata in try/except for robustness by @jeffreyftang in https://github.com/ludwig-ai/ludwig/pull/2373
- Reduce dropout prob in test_conv1d_stack by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2380
- fever: change broken download links by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2381
- Add default split config by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2379
- Fix CI: Skip failing ray GBM tests by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2391
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2389
- Triton ensemble export by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2251
- Fix: Random dataset splitting with 0.0 probability for optional validation or test sets. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2382
- Print final training report as tabulated text. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2383
- Add Ray 2.0 to CI by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2337
- add GBM configs to benchmarking by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2395
- Optional artifact logging for MLFlow by @ShreyaR in https://github.com/ludwig-ai/ludwig/pull/2255
- Simplify ludwig.benchmarking.benchmark API and add ludwig benchmark CLI by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2394
- rename kaggle_api_key to kaggle_key by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2384
- use new URL for yosemite dataset by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2385
- Encoder refactor V2 by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2370
- re-enable GBM tests after new lightgbm-ray release by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2393
- Added option to log artifact location while creating mlflow experiment by @ShreyaR in https://github.com/ludwig-ai/ludwig/pull/2397
- Treat dataset columns as object dtype during first pass of handle_missing_values by @jeffreyftang in https://github.com/ludwig-ai/ludwig/pull/2398
- fix: ParameterMetadata JSON serialization bug by @ksbrar in https://github.com/ludwig-ai/ludwig/pull/2399
- Adds registry to organize backward compatibility updates around versions and config sections by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2335
- Include split column in explanation df by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2405
- Fix AimCallback to model_name as Run.name by @alberttorosyan in https://github.com/ludwig-ai/ludwig/pull/2413
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2410
- Hotfix: features eligible for shared params hyperopt by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2417
- Nest FC Params in Decoder by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2400
- Hyperopt Backwards Compatibility by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2419
- Investigating test_resnet_block_layer intermittent test failure by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2414
- fix: Remove duplicate option from
cell_typefield schema by @ksbrar in https://github.com/ludwig-ai/ludwig/pull/2428 - Test for parameter updates in Ludwig Combiners - Part 3 by @jimthompson5802 in https://github.com/ludwig-ai/ludwig/pull/2332
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2430
- Hotfix: Proc column missing in output feature schema by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2435
- Nest hyperopt parameters into decoder object by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2436
- Fix: Make the twitter bots modeling example runnable by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2433
- Add MLG-ULB creditcard fraud dataset by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2425
- Bugfix: non-number inputs to GBM by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2418
- GBM: log intermediate progress by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2421
- Fix: Upgrade ludwig config before schema validation by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2441
- Log warning for calibration if validation set is trivially small by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2440
- Fixes calibration and adds example scripts by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2431
- Add medical no-show appointments dataset by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2387
- Added conditional check for UNK token insertion into category feature vocab by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2429
- Ensure synthetic dataset unit tests to clean up extra files. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2442
- Added feature specific parameter test for hyperopt by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2329
- Fixed version transformation to accept user configs without ludwig_version by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2424
- Fix mulitple partition predict by @magdyksaleh in https://github.com/ludwig-ai/ludwig/pull/2422
- Cache jsonschema validator to reduce memory pressure by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2444
- [tests] Added more explicit lifecycle management to Ray clusters during tests by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2447
- Fix: explicit keyword args for seaborn plot fn by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2454
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2453
- Extended hyperopt to support nested configuration block parameters by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2445
- Consolidate missing value strategy to only include bfill and ffill by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2457
- fix: Switched Learning Rate to NonNegativeFloat Field by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2446
- Support GitHub Codespaces by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2463
- Enh: quality-of-life improvements for
export_torchscriptby @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2459 - Disables
batch_size: autofor CPU-only training by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2455 - buxfix: triton model version as a string by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2461
- Updating images to Ray 2.0.0 and CUDA 11.3 by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2390
- Loss, Split, and Defaults Schema Additions by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2439
- More precise resource usage tracking by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2363
- Summarizing performance metrics and resource usage results by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2372
- [release-0.6] Cherry-pick bugfixes from upstream by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2471
- [release-0.6] Cherry-pick upstream commits by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2473
- [release-0.6] Cherry-pick upstream by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2476
- Cherry-pick backwards-compatibility fixes by @jeffreyftang in https://github.com/ludwig-ai/ludwig/pull/2487
- [cherry-pick] Fixed usage of checkpoints for AutoML in Ray 2.0 (#2485) by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2491
- fix: Automatically assign title to OneOfOptionsField (#2480) by @ksbrar in https://github.com/ludwig-ai/ludwig/pull/2492
- [cherry-pick] Fixed stratified splitting with Dask (#1883) by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2494
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2505
- AUTO: Enable hyperopt to be launched from a ray client by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2504
- [cherry-pick] Pin transformers < 4.22 until issues resolved (#2495) by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2510
- [cherry-pick] Fix flaky ray nightly image test (#2493) by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2511
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2513
- Add in-memory dataset size calculation to dataset statistics and hyperopt (#2509) by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2518
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2521
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2528
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2534
- Cherrypick: Cleanup: move to per-module loggers instead of the global logging object by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2539
- Update version to 0.6rc1. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2529
- Add resource isolation to 0.6 and fix merge conflicts by @magdyksaleh in https://github.com/ludwig-ai/ludwig/pull/2538
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2542
- More resource isolation cherrypicks by @magdyksaleh in https://github.com/ludwig-ai/ludwig/pull/2544
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2546
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2552
- Pin ray nightly version to avoid test failures related to TensorDType… by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2559
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2557
- Update version to 0.6. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2549
New Contributors
Congratulations to our new contributors!
- @Marvjowa made their first contribution in https://github.com/ludwig-ai/ludwig/pull/2236
- @Dennis-Rall made their first contribution in https://github.com/ludwig-ai/ludwig/pull/2192
- @abidwael made their first contribution in https://github.com/ludwig-ai/ludwig/pull/2263
- @noahlh made their first contribution in https://github.com/ludwig-ai/ludwig/pull/2284
- @jeffkinnison made their first contribution in https://github.com/ludwig-ai/ludwig/pull/2316
- @andife made their first contribution in https://github.com/ludwig-ai/ludwig/pull/2358
- @alberttorosyan made their first contribution in https://github.com/ludwig-ai/ludwig/pull/2413
Full Changelog: https://github.com/ludwig-ai/ludwig/compare/v0.5.3...v0.6
v0.6rc1
1 year agoWhat's Changed
- [release-0.6] Cherry-pick bugfixes from upstream by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2471
- [release-0.6] Cherry-pick upstream commits by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2473
- [release-0.6] Cherry-pick upstream by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2476
- Cherry-pick backwards-compatibility fixes by @jeffreyftang in https://github.com/ludwig-ai/ludwig/pull/2487
- [cherry-pick] Fixed usage of checkpoints for AutoML in Ray 2.0 (#2485) by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2491
- fix: Automatically assign title to OneOfOptionsField (#2480) by @ksbrar in https://github.com/ludwig-ai/ludwig/pull/2492
- [cherry-pick] Fixed stratified splitting with Dask (#1883) by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2494
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2505
- AUTO: Enable hyperopt to be launched from a ray client by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2504
- [cherry-pick] Pin transformers < 4.22 until issues resolved (#2495) by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2510
- [cherry-pick] Fix flaky ray nightly image test (#2493) by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2511
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2513
- Add in-memory dataset size calculation to dataset statistics and hyperopt (#2509) by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2518
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2521
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2528
- AUTO: by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2534
- Cherrypick: Cleanup: move to per-module loggers instead of the global logging object by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2539
- Update version to 0.6rc1. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2529
Full Changelog: https://github.com/ludwig-ai/ludwig/compare/v0.6.beta...v0.6rc1
v0.6.beta
1 year agoWhat's Changed
- Fix ray nightly import by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2196
- Restructured split config and added datetime splitting by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2132
- enh: Implements
InferenceModuleas a pipelined module with separate preprocessor, predictor, and postprocessor modules by @brightsparc in https://github.com/ludwig-ai/ludwig/pull/2105 - Explicitly pass data credentials when reading binary files from a RayBackend by @jeffreyftang in https://github.com/ludwig-ai/ludwig/pull/2198
- MlflowCallback: do not end run on_trainer_train_teardown by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2201
- Fail hyperopt with full import error when Ray not installed by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2203
- Make convert_predictions() backend-aware by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2200
- feat: MVP for explanations using Integrated Gradients from captum by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2205
- [Torchscript] Adds GPU-enabled input types for Vector and Timeseries by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2197
- feat: Added model type GBM (LightGBM tree learner), as an alternative to ECD by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2027
- [Torchscript] Parallelized Text/Sequence Preprocessing by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2206
- feat: Adding feature type shared parameter capability for hyperopt by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2133
- Bump up version to 0.6.dev. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2209
- Define
FloatOrAutoandIntegerOrAutoschema fields, and use them. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2219 - Define a dataclass for parameter metadata. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2218
- Add explicit handling for zero-length image byte buffers to avoid cryptic errors by @jeffreyftang in https://github.com/ludwig-ai/ludwig/pull/2210
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2231
- Create dataset util to form repeatable train/vali/test split by @amholler in https://github.com/ludwig-ai/ludwig/pull/2159
- Bug fix: Use safe rename which works across filesystems when writing checkpoints by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2225
- Add parameter metadata to the trainer schema. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2224
- Add an explicit call to merge_wtih_defaults() when loading a config from a model directory. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2226
- Fixes flaky test test_datetime_split[dask] by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2232
- Fixes prediction saving for models with Set output by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2211
- Make ExpectedImpact JSON serializable by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2233
- standardised quotation marks, added missing word by @Marvjowa in https://github.com/ludwig-ai/ludwig/pull/2236
- Add boolean postprocessing to dataset type inference for automl by @magdyksaleh in https://github.com/ludwig-ai/ludwig/pull/2193
- Update get_repeatable_train_val_test_split to handle non-stratified split w/ no existing split by @amholler in https://github.com/ludwig-ai/ludwig/pull/2237
- Update R2 score to handle single sample computation by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2235
- Input/Output Feature Schema Refactor by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2147
- Fix nan in entmax loss and flaky sparsemax/entmax loss tests by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2238
- Fix preprocessing dataset split API backwards compatibility upgrade bug. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2239
- Removing duplicates in constants from recent PRs by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2240
- Add attention scores of the vit encoder as an additional return value by @Dennis-Rall in https://github.com/ludwig-ai/ludwig/pull/2192
- Unnest Audio Feature Preprocessing Config by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2242
- Fixed handling of invalud number values to treat as missing values by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2247
- Support saving numpy predictions to remote FS by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2245
- Use global constant for description.json by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2246
- Removed import warnings when LightGBM and Ray not requested by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2249
- Adds ability to read images from numpy files and numpy arrays by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2212
- Hyperopt steps per epoch not being computed correctly by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2175
- Fixed splitting when providing pre-split inputs by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2248
- Added Backwards Compatibility for Audio Feature Preprocessing by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2254
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2256
- Fix: Don't skip saving the model if the save path already exists. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2264
- Load best weights outside of finally block, since load may throw an exception by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2268
- Reduce number of distributed tests. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2270
- [WIP] Adds
inference_utils.pyby @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2213 - Run github checks for pushes and merges to *-stable. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2266
- Add ludwig logo and version to CLI help text. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2258
- Add hyperopt_statistics.json constant by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2276
- fix: Make
BaseTrainerConfigan abstract class by @ksbrar in https://github.com/ludwig-ai/ludwig/pull/2273 - [Torchscript] Adds
--deviceargument toexport_torchscriptCLI command by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2275 - Use pytest tmpdir fixture wherever temporary directories are used in tests. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2274
- adding configs used in benchmarking by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2263
- Fixes #2279 by @noahlh in https://github.com/ludwig-ai/ludwig/pull/2284
- adding hardware usage and software packages tracker by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2195
- benchmarking utils by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2260
- dataclasses for summarizing benchmarking results by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2261
- Benchmarking core by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2262
- Fixed default eval_batch_size when setting batch_size=auto by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2286
- Remove obsolete postprocess_inference_graph function. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2267
- [Torchscript] Adds BERT tokenizer + partial HF tokenizer support by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2272
- Support passing ground_truth as df for visualizations by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2281
- catching urllib3 exception by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2294
- Run pytest workflow on release branches. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2291
- Save checkpoint if train_steps is smaller than batcher's steps_per_epoch by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2298
- Fix typo in amazon review datasets: s/review_tile/review_title by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2300
- Refactor non-distributed automl utils into a separate directory. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2296
- Don't skip normalization in TabNet during inference on a single row. by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2299
- Fix error in postproc_predictions calculation in model.evaluate() by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2304
- Test for parameter updates in Ludwig components by @jimthompson5802 in https://github.com/ludwig-ai/ludwig/pull/2194
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2311
- Use warnings to suppress repeated logs for failed image reads by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2312
- Use ray dataset and drop type casting in binary_feature prediction post processing for speedup by @magdyksaleh in https://github.com/ludwig-ai/ludwig/pull/2293
- Add size_bytes to DatasetInfo and DataSource by @jeffreyftang in https://github.com/ludwig-ai/ludwig/pull/2306
- Fixes TensorDtype TypeError in Ray nightly by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2320
- Add configuration section for global feature parameters by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2208
- Ensures unit tests are deleting artifacts during teardown by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2310
- Fixes unit test that had empty Dask partitions after splitting by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2313
- Serve json numpy encoding by @jeffkinnison in https://github.com/ludwig-ai/ludwig/pull/2316
- fix: Mlflow config being injected in hyperopt config by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2321
- Update tests that use preprocessing to match new defaults config structure by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2323
- Bump test timeout to 60 minutes by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2325
- Set a default value for size_bytes in DatasetInfo by @jeffreyftang in https://github.com/ludwig-ai/ludwig/pull/2331
- Pin nightly versions to fix CI by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2327
- Log number of failed image reads by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2317
- Add test with encoder dependencies for global defaults by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2342
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2334
- Add wine quality notebook to demonstrate using config defaults by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2333
- fix: GBM tests failing after new release from upstream dependency by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2347
- fix: restore overwrite of eval_batch_size on GBM schema by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2345
- Removes empty partitions after dropping rows and splitting datasets by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2328
- fix: Properly serialize
ParameterMetadatato JSON by @ksbrar in https://github.com/ludwig-ai/ludwig/pull/2348 - Test for parameter updates in Ludwig Components - Part 2 by @jimthompson5802 in https://github.com/ludwig-ai/ludwig/pull/2252
- refactor: Replace bespoke marshmallow fields that accept multiple types with a new 'combinatorial'
OneOfFieldthat accepts other fields as arguments. by @ksbrar in https://github.com/ludwig-ai/ludwig/pull/2285 - Use Ray Datasets to read binary files in parallel by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2241
- typos: Update README.md by @andife in https://github.com/ludwig-ai/ludwig/pull/2358
- Respect the resource requests in RayPredictor by @magdyksaleh in https://github.com/ludwig-ai/ludwig/pull/2359
- Resource tracker threading by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2352
- Allow writing init_config results to remote filesystems by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2364
- Fixed export_mlflow command to not assume an existing registered_model_name by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2369
- fix: Fixes to serialization, and update to allow set repo location. by @brightsparc in https://github.com/ludwig-ai/ludwig/pull/2367
- Add amazon employee access challenge kaggle dataset by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2349
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2362
- Wrap read of cached training set metadata in try/except for robustness by @jeffreyftang in https://github.com/ludwig-ai/ludwig/pull/2373
- Reduce dropout prob in test_conv1d_stack by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2380
- fever: change broken download links by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2381
- Add default split config by @hungcs in https://github.com/ludwig-ai/ludwig/pull/2379
- Fix CI: Skip failing ray GBM tests by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2391
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2389
- Triton ensemble export by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2251
- Fix: Random dataset splitting with 0.0 probability for optional validation or test sets. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2382
- Print final training report as tabulated text. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2383
- Add Ray 2.0 to CI by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2337
- add GBM configs to benchmarking by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2395
- Optional artifact logging for MLFlow by @ShreyaR in https://github.com/ludwig-ai/ludwig/pull/2255
- Simplify ludwig.benchmarking.benchmark API and add ludwig benchmark CLI by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2394
- rename kaggle_api_key to kaggle_key by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2384
- use new URL for yosemite dataset by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2385
- Encoder refactor V2 by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2370
- re-enable GBM tests after new lightgbm-ray release by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2393
- Added option to log artifact location while creating mlflow experiment by @ShreyaR in https://github.com/ludwig-ai/ludwig/pull/2397
- Treat dataset columns as object dtype during first pass of handle_missing_values by @jeffreyftang in https://github.com/ludwig-ai/ludwig/pull/2398
- fix: ParameterMetadata JSON serialization bug by @ksbrar in https://github.com/ludwig-ai/ludwig/pull/2399
- Adds registry to organize backward compatibility updates around versions and config sections by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2335
- Include split column in explanation df by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2405
- Fix AimCallback to model_name as Run.name by @alberttorosyan in https://github.com/ludwig-ai/ludwig/pull/2413
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2410
- Hotfix: features eligible for shared params hyperopt by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2417
- Nest FC Params in Decoder by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2400
- Hyperopt Backwards Compatibility by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2419
- Investigating test_resnet_block_layer intermittent test failure by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2414
- fix: Remove duplicate option from
cell_typefield schema by @ksbrar in https://github.com/ludwig-ai/ludwig/pull/2428 - Test for parameter updates in Ludwig Combiners - Part 3 by @jimthompson5802 in https://github.com/ludwig-ai/ludwig/pull/2332
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2430
- Hotfix: Proc column missing in output feature schema by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2435
- Nest hyperopt parameters into decoder object by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2436
- Fix: Make the twitter bots modeling example runnable by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2433
- Add MLG-ULB creditcard fraud dataset by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2425
- Bugfix: non-number inputs to GBM by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2418
- GBM: log intermediate progress by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2421
- Fix: Upgrade ludwig config before schema validation by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2441
- Log warning for calibration if validation set is trivially small by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2440
- Fixes calibration and adds example scripts by @dantreiman in https://github.com/ludwig-ai/ludwig/pull/2431
- Add medical no-show appointments dataset by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2387
- Added conditional check for UNK token insertion into category feature vocab by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2429
- Ensure synthetic dataset unit tests to clean up extra files. by @justinxzhao in https://github.com/ludwig-ai/ludwig/pull/2442
- Added feature specific parameter test for hyperopt by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2329
- Fixed version transformation to accept user configs without ludwig_version by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2424
- Fix mulitple partition predict by @magdyksaleh in https://github.com/ludwig-ai/ludwig/pull/2422
- Cache jsonschema validator to reduce memory pressure by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2444
- [tests] Added more explicit lifecycle management to Ray clusters during tests by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2447
- Fix: explicit keyword args for seaborn plot fn by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2454
- [pre-commit.ci] pre-commit suggestions by @pre-commit-ci in https://github.com/ludwig-ai/ludwig/pull/2453
- Extended hyperopt to support nested configuration block parameters by @tgaddair in https://github.com/ludwig-ai/ludwig/pull/2445
- Consolidate missing value strategy to only include bfill and ffill by @arnavgarg1 in https://github.com/ludwig-ai/ludwig/pull/2457
- fix: Switched Learning Rate to NonNegativeFloat Field by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2446
- Support GitHub Codespaces by @jppgks in https://github.com/ludwig-ai/ludwig/pull/2463
- Enh: quality-of-life improvements for
export_torchscriptby @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2459 - Disables
batch_size: autofor CPU-only training by @geoffreyangus in https://github.com/ludwig-ai/ludwig/pull/2455 - buxfix: triton model version as a string by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2461
- Updating images to Ray 2.0.0 and CUDA 11.3 by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2390
- Loss, Split, and Defaults Schema Additions by @connor-mccorm in https://github.com/ludwig-ai/ludwig/pull/2439
- More precise resource usage tracking by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2363
- Summarizing performance metrics and resource usage results by @abidwael in https://github.com/ludwig-ai/ludwig/pull/2372
New Contributors
- @Marvjowa made their first contribution in https://github.com/ludwig-ai/ludwig/pull/2236
- @Dennis-Rall made their first contribution in https://github.com/ludwig-ai/ludwig/pull/2192
- @abidwael made their first contribution in https://github.com/ludwig-ai/ludwig/pull/2263
- @noahlh made their first contribution in https://github.com/ludwig-ai/ludwig/pull/2284
- @jeffkinnison made their first contribution in https://github.com/ludwig-ai/ludwig/pull/2316
- @andife made their first contribution in https://github.com/ludwig-ai/ludwig/pull/2358
- @alberttorosyan made their first contribution in https://github.com/ludwig-ai/ludwig/pull/2413
Full Changelog: https://github.com/ludwig-ai/ludwig/compare/v0.5.3...v0.6.beta